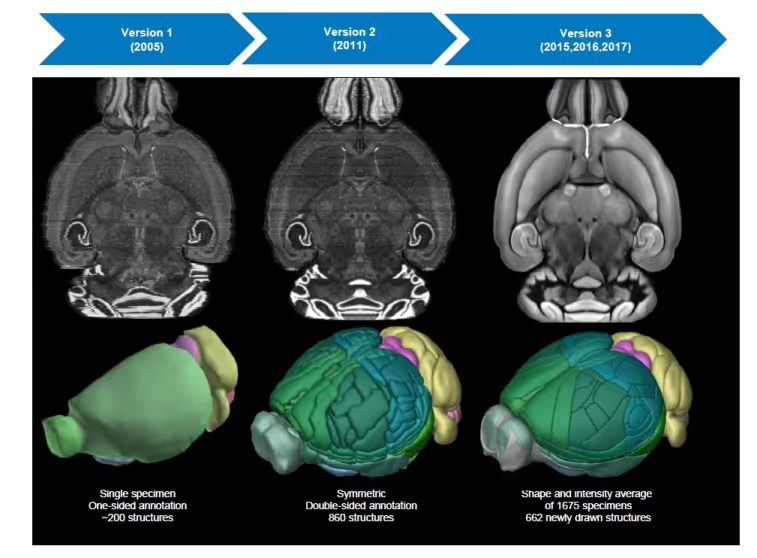
Common Coordinate Frameworks for the BICCN
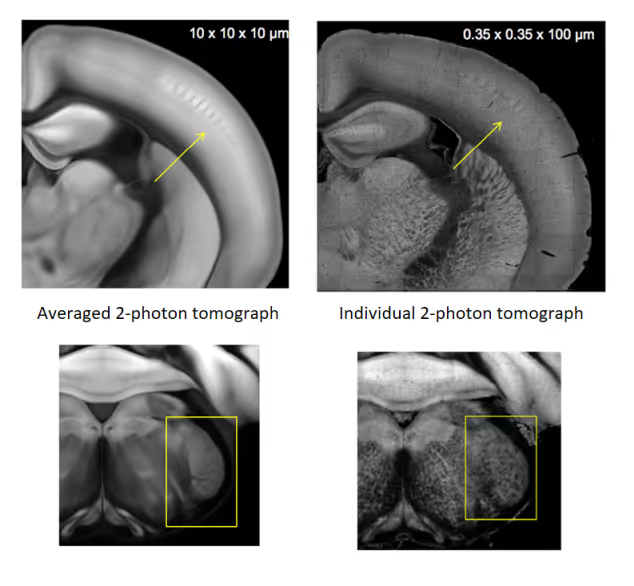
Spatial Data Management
BICCN Standards, Best Practices, and Recommendations
Metadata and File Formats
High level descriptive metadata to describe BICCN data collections. Instructions and templates for providing information about projects and data collections, and their descriptive metadata which must be submitted for each data collection milestone. In use by: BCDC for the BICCN Data Catalog.
Basic descriptive metadata for samples and specimens profiled in BICCN. Instructions and templates for data submission are provided. An inventory of samples and associated cells or brains profiled for each sample for each dataset, reported on a quarterly basis. This is suitable for identifying the transgenic line and anatomic region sampled in a manageable spreadsheet, but is not intended to include more detailed provenance. In use by: BCDC for the BICCN Data Catalog.
Metadata standard for a 3D microscopy dataset to support reuse by scientists who did not generate the data. 3D microscopy data submitted to BIL must adhere to this metadata standard. In use by: BIL Data Archive.
NeuroData Without Borders (NWB2.0)
Data standard for neurophysiology, providing neuroscientists with a common standard to share, archive, use, and build common analysis tools for neurophysiology data. In use by: DANDI Archive.Endorsed by: INCF.
Data standard for neuroimaging data (e.g., MRI). In use by: DANDI Archive.
A proposal extending Brain Imaging Data Structure (BIDS) to 3D Microscopy. 3D microscopy submitted specifically to DANDI archive will adhere to this standard. In use by: DANDI Archive.
Open data format for encoding neuro morphologies. Required for morphological data submitted to BIL and NeuroMorpho.org. NeuroMorpho.org uses the tool L-Measure to convert common formats into SWC. In use by: BIL Data Archive, Neuromorpho.org.
NeuroMorpho documentation on use of SWC
Discussion on SWC at the NEURON forum
Spatial and Semantic Standards
The Allen Mouse Common Coordinate Framework (CCF) is the main anatomic data browser and coordinate environment for mouse data within the BICCN. If you are acquiring spatial data in the mouse, it should be registered to the CCF V3. Registered data is available through BIL data archive. Anatomic terms associated with the this CCF are used as annotations of samples and specimens. In use by: BCDC for the BICCN Data Catalog.
List of tools for registering images.
Templates for registering different data types.
The Allen Human Atlas - 3D is a is a parcellation of the non-linear average of the MNI152 using the same anatomical ontology as the section based Allen Human Reference Atlas. If you are acquiring samples in the human, approximate location of the sample should be registered to the Allen Human Atlas - 3D. In use by: BCDC for the BICCN Data Catalog.
Ontology for describing electrophysiology stimulation parameters, developed by an INCF cross project working group. This ontology is suitable for use in NWB formats.
Ontology for multi-species anatomical terms. All anatomical metadata should be mapped to UBERON identifiers or a particular atlas. A mapping between the Allen Brain Atlas has been completed. If the latter, the full reference for the atlas should be supplied.
Covers major neuroscience techniques. Based on NIFSTD methods ontology, OBI and EFO. Use this ontology to map techniques, modalities, assays, devices and tools to a controlled vocabulary. In use by: BCDC for the BICCN Data Catalog, BIL, NeMO
Instructions on working with and adding a term can be found here.
Ontology for representing data-defined cell types (omics) from the BICCN flagship paper. Used by BRAIN Data Standards group for -omics defined cell types. In use by: BICCN Mouse Primary Cortex Mini-Atlas – Cell Types Explorer.
Identifiers and Protocols
ORCIDs are unique identifiers for researchers. All investigators should provide ORCIDs for contributors to datasets to facilitate cross-dataset linkage and to receive credit for publishing datasets.
See documentation at individual archives for adding ORCIDs.
Unique identifier for antibodies, digital tools, cell lines, organisms, plasmids and biosamples. If you create a new tool for BICCN, please obtain an RRID. In use by: BICCN Data Catalog, BICCN.org, journals and publications.
BICCN investigators are required to share experimental protocols. Protocols.io is one option for this that provides a DOI and version history for protocols. In use by: BICCN members, BICCN Data Catalog
BICCN Working Groups
BICCN 2.0 Whole Mouse Brain WG. Chairs: Eran Mukamel, UCSD, Hongkui Zeng, Allen Institute
- Data integration for transcriptomic & epigenomic data across entire mouse brain and spinal cord
- Pipeline processing
- Collaborative analysis to support the creation of a cell census and consortium publication(s)
- Support FAIR data standards
- Create high impact outputs of the scientific work
- Omics standards
- Sample level QC
- Sequence data mapping
- Cell level standards
- Cluster level standards
- Uniform pipelines and BICCN Whole Mouse Brain Workspace
- Terra workspace
BICCN 2.0 Cell-Type Specific Enhancers. Chairs: Bosiljka Tasic, Allen Institute, Bing Ren, UCSD
- Develop methods to identify cell-type specific enhancers that can drive systemic delivery of reporter genes to select subclasses or types of brain cells in mice and primates.
- Obtain experimental data to support the validity of these methods.
- Single-cell multiomic atlases of the primary motor cortex in mice and primates
- Computational tools for predicting brain cell-type specific enhancers
- Experimentally validated enhancers active in select cortical neuronal sub-types
BICCN Infrastructure & Metadata WG. Chairs: Carol Thompson, Allen Institute, Mike Hawrylycz, Allen Institute
- Data standards
- Data collection processes and reporting
- BICCN infrastructure ecosystem improvements
- Support FAIR data standards
- Collection level metadata
- Specimen metadata
- Harmonized metadata across BCDC and archives
- Ontologies
- Neuroscience methods ontology
BICCN Morphology WG. Chairs: Giorgio Ascoli, GMU, Hanchuan Peng, SEU-Allen Joint Center
- Morphology metadata
- Data formats
- Standards
- CCF mapping
- Reconstructions
- Community usage
- Standardized on NIFTI and SWC data formats
- Standard metadata file (metainfo.txt)
- fMOST QC checks via Data Core.
- Data quality
- Reconstruction difficulty assessment
- Reference annotation and reference terminologies
- CCF 3.0 visualization
- JSON format for soma locations and projection targets
- Standardized inventory of BICCN cell morphologies
BICCN 2.0 Molecular Wiring Diagrams Working Group WG. Chair: Hongwei Dong
- Begin planning on BICCN 2.0 project on Molecularly Annotated Wiring Diagrams of the Mammalian Brain
- Finalize the mini-atlas joint anatomy paper
BICCN Mini-Atlas Flagship Publication WG. Chairs: Joe Ecker, Salk Institute, Hongkui Zeng, Allen Institute, John Ngai, NIH
- Planning, coordinating and writing the mini-atlas flagship paper.
- Supporting FAIR data standards in the mini-atlas paper and across companion papers.
Complete
BICCN Human & NHP WG. Chairs: Ed Lein, Allen Institute, Arnold Kriegstein, UCSF
- Coordination on regional sampling plans
- Methods comparisons
- Scope out future big human and NHP atlasing
- Joint comparative analysis
Active
BICCN 2.0 Developing Brain Atlas WG. Chairs: Paola Arlotta, Harvard University, Tomasz Nowakowski, UCSF
- Mouse, human, and NHP development
Active
BICCN 2.0 Proteomic Brain Atlas WG. Chairs: Kwanghun Chung, MIT and Elizabeth Hillman, Columbia University
- Technology platform development
- Proteomic data acquisition and integration
- Collaborative analysis
- Technology, reagents, protocols, and data dissemination
- Formats: BIDS and BIDS microscopy extension
- Compression and serializable access
- Standardized metadata
- Protocols
- Sample/organisms
- Behavioral/clinical
- Resolution
Toward an Integrated BICCN Data Set (IDS)
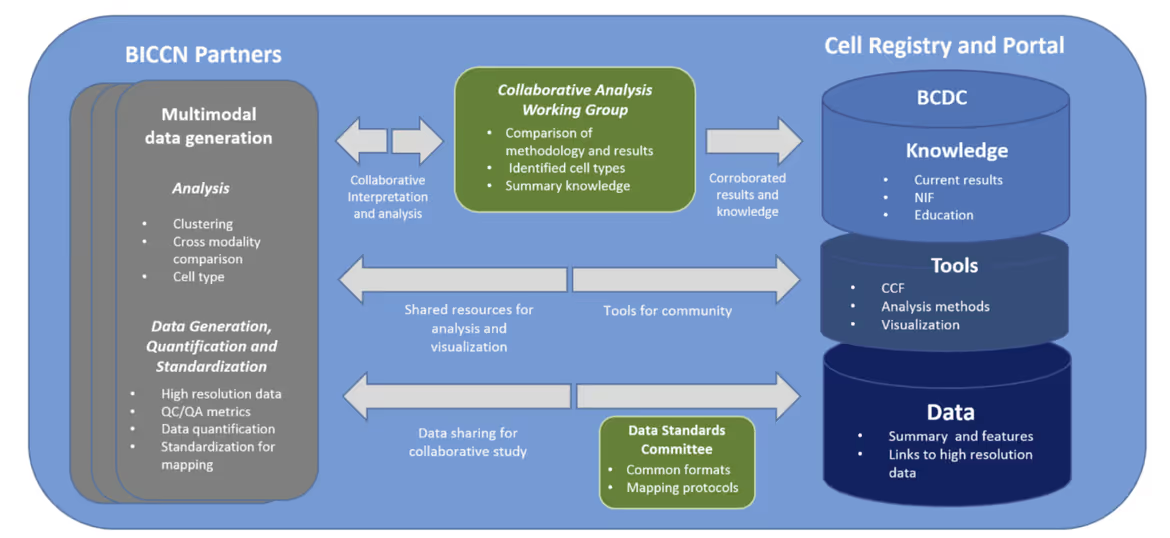
Data, Tools, and Knowledge
Data Levels
- Level 0: (Raw) Raw data including mapped sequence reads, raw non- or minimally-compressed image series. Multivariate data sets are generated where applicable. These data are center specific and not meant for being publicly distributed or distributed to the BCDC. a div block.
- Level 1: (QA/QC) Preprocessed data sets having passed an initial round of QA/QC or computation. This is data intended for sharing within the BICCN network and storage in R24 archive centers, and it uses the appropriate format and compression level for derived computation and analysis.
- Level 2: (Linked) These data are mapped to a spatially determined region or stereotaxic site. Ultimately this data is accessible and displayable within the Common Coordinate Framework (CCF.)
- Level 3: (Featured) A derived data set summarizing relevant features and properties for a modality of interest. This level enables shared and novel algorithmic approaches across or within centers. Examples include differential expression, clustering analyses, cross modality analysis.
- Level 4: (Integrated) Informative data sets and that meet standards for public deployment and have value for the BICCN network and larger community in advancing our understanding of cell types in the brain. Appropriate results will be release on the BICCN portal.
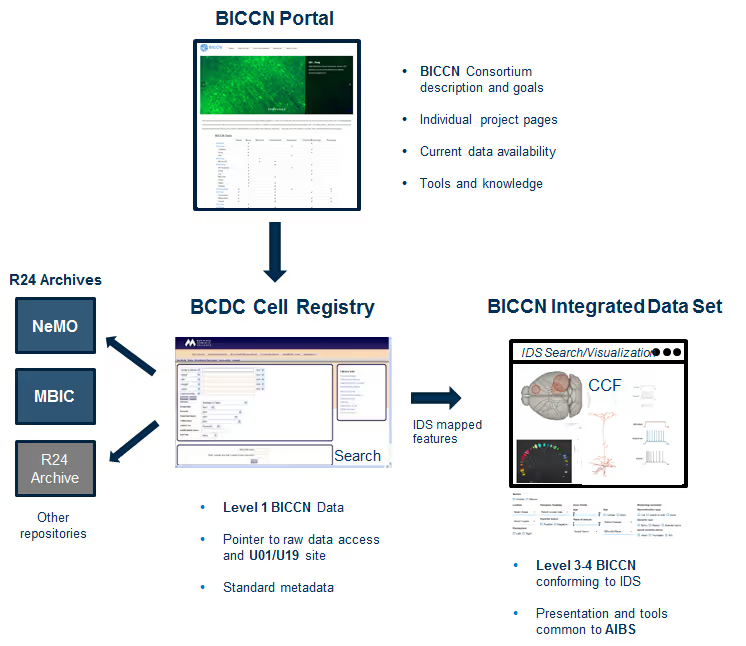
The BCDC Cell Registry and BICCN Integrated Data Set