The Allen Human Reference Atlas – 3D, 2020 is a 3-dimensional annotated parcellation of the adult human brain. The atlas includes 141 brain regions spanning the complete volume of the MRI reference brain “ICBM 2009b Nonlinear Symmetric”, a non-linear average of the MNI152 database of 152 normal brain images. These structures include regions from a 2D plate-based histological reference atlas of the adult human brain (Ding et al., 2016), that can be identified in the average MRI volume. The atlas is intended to serve as a positional common coordinate framework for mapping adult human brain data generated across the BICCN. (Allen Human Reference Atlas, 3D, 2020; RRID:SCR_017764)
Access the Allen Human Reference Atlas - 3D, 2020 here:
https://community.brain-map.org/t/allen-human-reference-atlas-3d-2020-new/405
Allen Institute, Brain Science (https://alleninstitute.org/what-we-do/brain-science/)
Enhanced and Unified Anatomical Labeling for a Common Mouse Brain Atlas - To facilitate comparison between existing atlases, the Franklin and Paxinos (FP) label was imported and refined into the Allen Common Coordinate Framework (CCF). Cell type specific transgenic mice and an MRI atlas were used to adjust and further segment the labels. Moreover, new segmentations were created in the dorsal striatum using cortico-striatal connectivity data. The anatomical labels were digitized based on the Allen ontology, and a web-interface was created for easy visualization. These labels provide a resource to isolate and identify mouse brain anatomical structures.
Access anatomical labels here:
http://kimlab.io/brain-map/atlas/
Yongsoo Kim lab (http://kimlab.io/), Penn State University
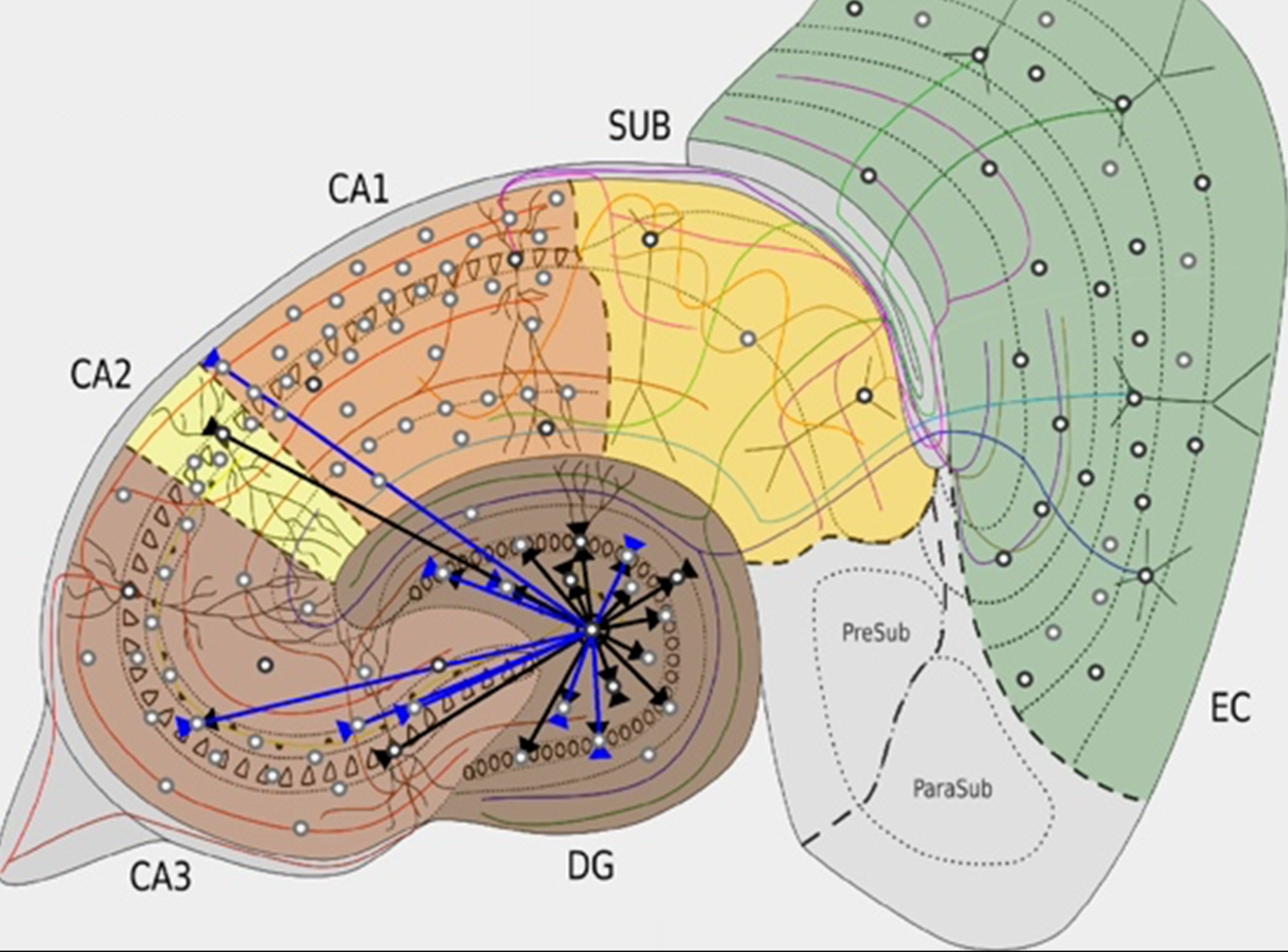
Hippocampome.org is a curated knowledge base of the neuron types of the rodent hippocampal formation. Knowledge concerning the morphology, electrophysiology, molecular expression, and connectivity of cells in the dentate gyrus, CA3, CA2, CA1, subiculum, and entorhinal cortex is distilled from published evidence and is continuously updated as new information becomes available. Each reported neuronal property is documented with a pointer to, and excerpt from, relevant published evidence, such as citation quotes or illustrations. (Hippocampome.org, RRID:SCR_009023)
Access Hippocampome here:
http://hippocampome.org/php/index.php
Giorgio Ascoli lab (https://krasnow1.gmu.edu/cn3/ascoli/), George Mason University
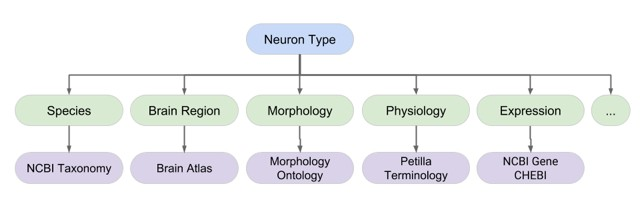
Neuron Phenotype Ontology - An important aspect in the classification of cell type identity and function is the use of powerful and flexible ontologies. The Neuron Phenotype Ontology is a prototype system for managing neuronal phenotypes that spans across the different phases of knowledge discovery. The system comprises a knowledge base of neuron types and supporting python codes, and supports the concepts of common usage types and evidence based models. It enables researchers to generate a complex neuron phenotype out of any number of individual phenotypes. Phenotype values are tied to formal biomedical ontologies, ensuring a consistent semantic representation and that the phenotypes themselves can be integrated with other types of data. Version 1.0 of the ontology is currently available. (NIFSTD, RRID:SCR_005414)
Access the Neuron Phenotype Ontology here:
Neuron Phenotype Ontology: https://github.com/SciCrunch/NIF-Ontology/blob/master/docs/Neurons.md
IPython notebook for generating your own neuron names: https://github.com/tgbugs/pyontutils/blob/master/neurondm/docs/NeuronLangExample.ipynb
Neuroscience Information Framework (https://neuinfo.org/), University of California, San Diego
BICCN Pipelines
Overview
Pipeline Standards, Maintenance, and Availability
- Open-access and developed with GA4GH standards.
- Written in the Workflow Description Language (WDL), a community-maintained, human-readable workflow language that can run on Cromwell, a portable execution engine that can be launched anywhere, locally or in the cloud.
- Containerized using public Docker instances, allowing researchers to exactly reproduce the workflow software.
- Code is developed and maintained in the WDL Analysis Research Pipelines (WARP) repository in GitHub. Overviews and workflow code for BICCN-related pipelines are linked in the table above; additionally, relevant pipelines can be identified by typing the keyword “BICCN” in the WARP Documentation search bar.The WARP Overview details navigating the repository, pipeline development, and running the workflows.
- Workflows are available for export from Dockstore, a GA4GH-compliant platform for sharing Docker-based tools. Search “warp” on Dockstore to find all WARP pipelines, including those used in the BICCN.
- Workflows are also available to test on Terra, the cloud-based bioinformatics platform used for BCDC data processing. To get started, register for Terra using the registration guide. To try a pipeline, navigate to the pipeline’s workspace linked in the table above or search for the “BICCN” tag in the workspaces tag search bar. Each workspace contains downsampled data, detailed instructions for using the workflows, and cost guidelines. Learn more about Terra with the Getting Started guides.
Citing the Pipelines
Additional Single-cell Transcriptomic Pipeline Resources
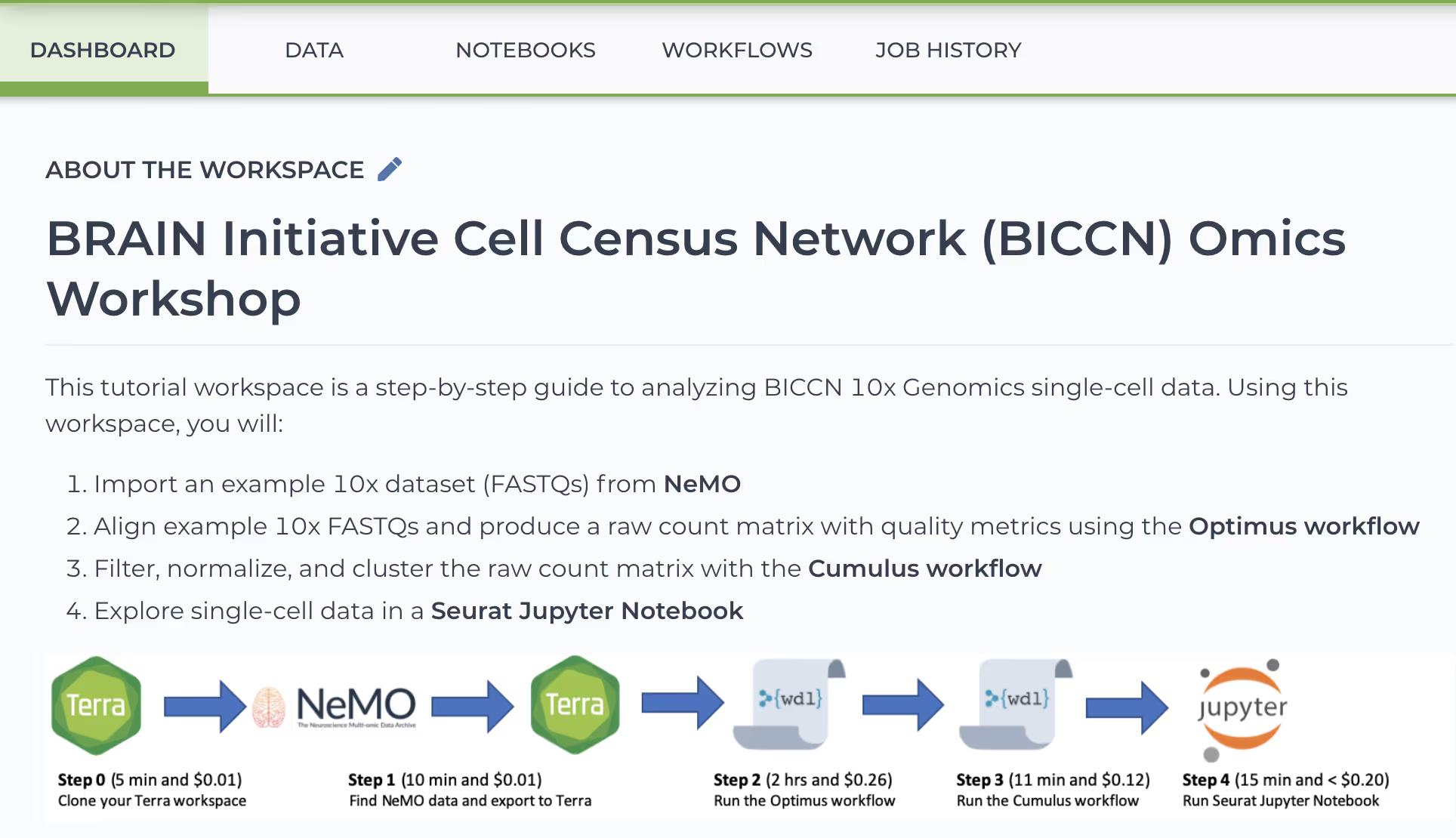
BICCN Omics Workshop Workspace
- Import an example 10x dataset (FASTQs) from NeMO
- Align example 10x FASTQs and produce a raw count matrix with quality metrics using the Optimus workflow
- Filter, normalize, and cluster the raw count matrix with the Cumulus workflow
- Explore single-cell data in a Seurat Jupyter Notebook