QuickNII - A major challenge in data mapping is the registration and alignment of 2D image sections in a 3D context. QuickNII is a stand-alone software tool for semi-automated affine spatial registration of sectional image data to a 3D reference atlas coordinate framework and can interact with the Allen Institute Common Coordinate Framework. A key feature in the tool is the capability to generate user defined cut planes through the reference atlas by matching the orientation of the cut plane of the sectional image data. The reference atlas is transformed to match anatomical landmarks in the corresponding experimental images. In this way, the spatial relationship between experimental image and atlas is defined, without introducing distortions in the original experimental images. (QuickNII, RRID:SCR_016854)
Access QuickNII here:
https://www.nitrc.org/projects/quicknii/
Inquiries and support: support@humanbrainproject.eu
J.G. Bjaalie / Neural Systems Lab (https://www.med.uio.no/imb/english/research/groups/neural-systems/), University of Oslo
Human Brain Project (http://www.humanbrainproject.eu/)
HistoloZee 2.0 is a software tool for interactively mapping 2D and 3D molecular and anatomical histology into Common Coordinate Frameworks (CCFs). It has simple, interactive registration workflows that connect user images with CCFs. Planned features include:
- Concurrent rendering of high-resolution histology and 3D images
- Slicing of images at arbitrary orientations
- Seamless multi-scale zooming
- Surface meshes of anatomical structures
- Interactive affine registration
- User-guided deformable registration
- Persistence of projects in human-readable format
- Loading of common histology and medical image formats
Access HistoloZee 2.0 here:
http://picsl.upenn.edu/software/histolozee/
Penn Image Computing and Science Lab (PICSL) (http://picsl.upenn.edu/software/histolozee/), University of Pennsylvania
Cell Locator is a tool for placing tissue samples into annotated 3D coordinate systems. Users can draw polylines onto arbitrarily oriented slices of the 3D space, give them a physical thickness, and save the results to the JSON file.
This tool is intended to be used by members of the BICCN to localize their cells and tissue samples so that data can be described in a consistent spatial framework. The tool currently supports the Allen Common Coordinate Framework v3, and will be extended to support a human reference space soon.
Cell Locator was developed by Kitware, Inc. It currently supports Windows, and will soon have installers for MacOS and Linux.
Access Cell Locator here:
https://github.com/BICCN/cell-locator/
Allen Institute for Brain Science (https://alleninstitute.org/what-we-do/brain-science/)
Kitware, Inc. (https://www.kitware.com/)
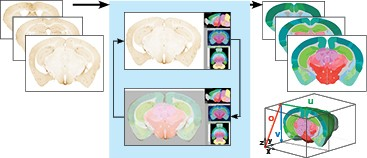
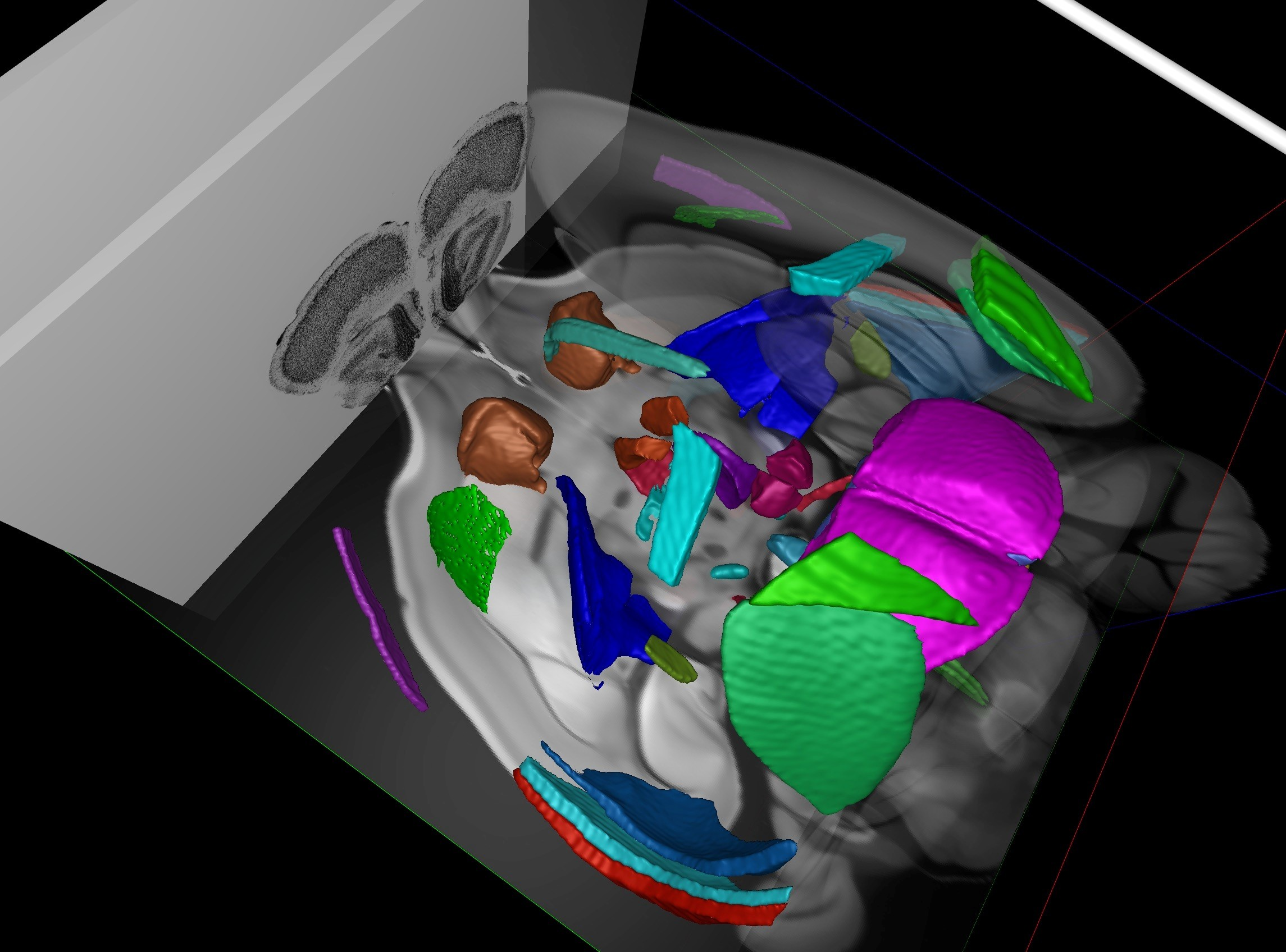
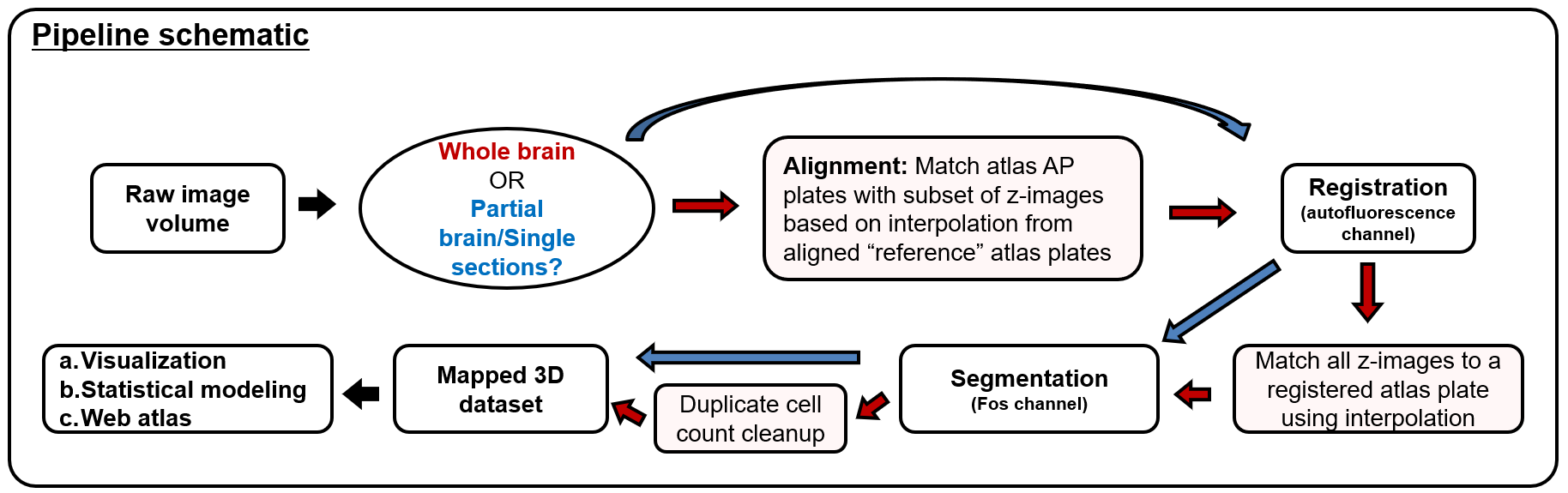
SMART (semi-manual alignment to reference templates) extends the WholeBrain framework in R for segmenting and registering experimental images to the Allen Mouse Common Coordinate Framework (CCF). SMART streamlines the processing of large volumetric LSFM datasets and solves issues with non-uniform morphing across the anterior-posterior axis with an interactive “choice game.” It also accounts for duplicate cell counts in adjacent z images and presents new ways to easily parse apart and interactively visualize the final mapped datasets.
Behavioral Neuroscience Research Branch, The National Institute on Drug Abuse Intramural Research Program (https://irp.drugabuse.gov/organization/bnrb/)
BICCN Pipelines
Overview
Pipeline Standards, Maintenance, and Availability
- Open-access and developed with GA4GH standards.
- Written in the Workflow Description Language (WDL), a community-maintained, human-readable workflow language that can run on Cromwell, a portable execution engine that can be launched anywhere, locally or in the cloud.
- Containerized using public Docker instances, allowing researchers to exactly reproduce the workflow software.
- Code is developed and maintained in the WDL Analysis Research Pipelines (WARP) repository in GitHub. Overviews and workflow code for BICCN-related pipelines are linked in the table above; additionally, relevant pipelines can be identified by typing the keyword “BICCN” in the WARP Documentation search bar.The WARP Overview details navigating the repository, pipeline development, and running the workflows.
- Workflows are available for export from Dockstore, a GA4GH-compliant platform for sharing Docker-based tools. Search “warp” on Dockstore to find all WARP pipelines, including those used in the BICCN.
- Workflows are also available to test on Terra, the cloud-based bioinformatics platform used for BCDC data processing. To get started, register for Terra using the registration guide. To try a pipeline, navigate to the pipeline’s workspace linked in the table above or search for the “BICCN” tag in the workspaces tag search bar. Each workspace contains downsampled data, detailed instructions for using the workflows, and cost guidelines. Learn more about Terra with the Getting Started guides.
Citing the Pipelines
Additional Single-cell Transcriptomic Pipeline Resources
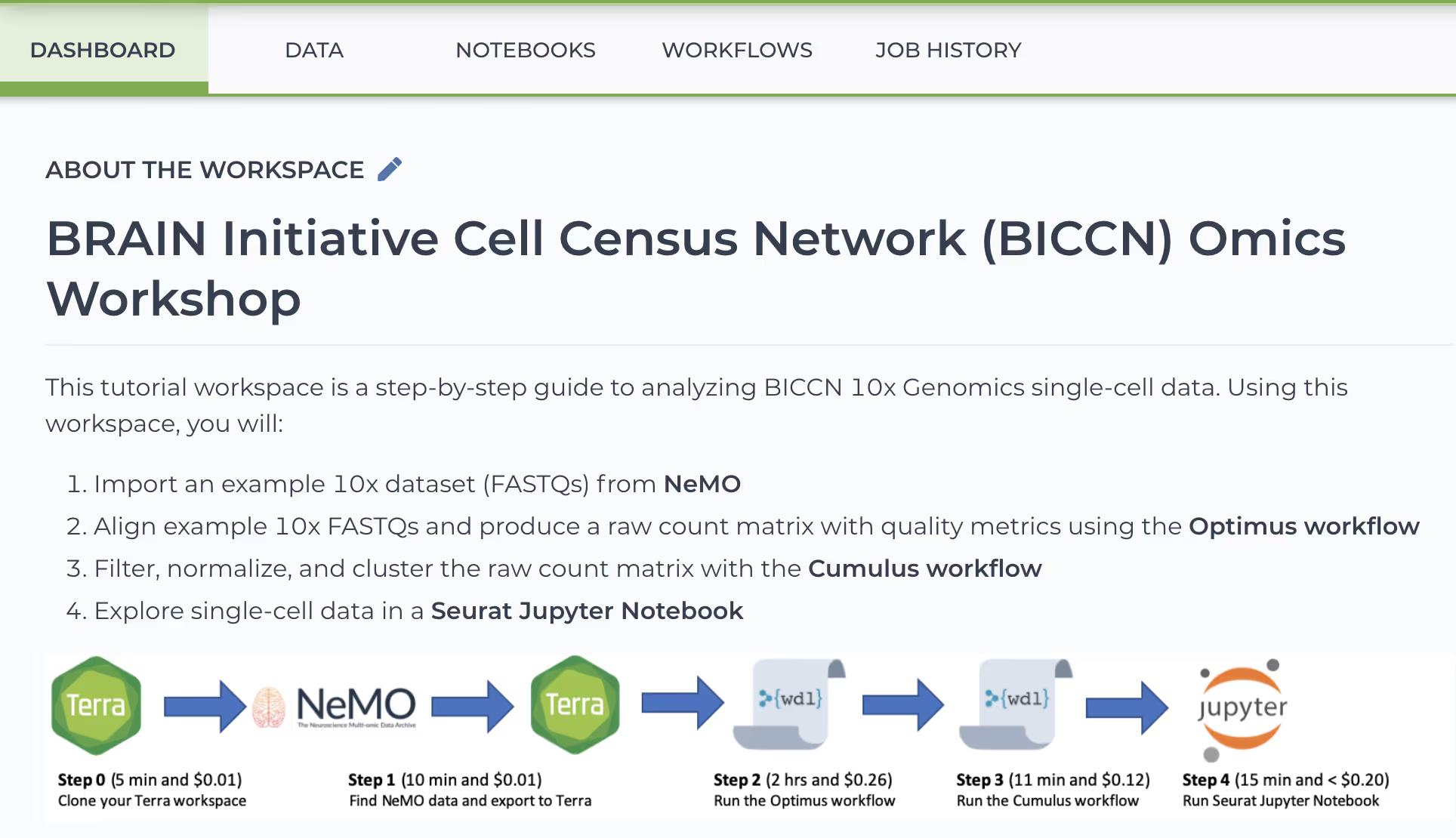
BICCN Omics Workshop Workspace
- Import an example 10x dataset (FASTQs) from NeMO
- Align example 10x FASTQs and produce a raw count matrix with quality metrics using the Optimus workflow
- Filter, normalize, and cluster the raw count matrix with the Cumulus workflow
- Explore single-cell data in a Seurat Jupyter Notebook