Linking molecular and anatomical features of brain cell identity through computational data integration
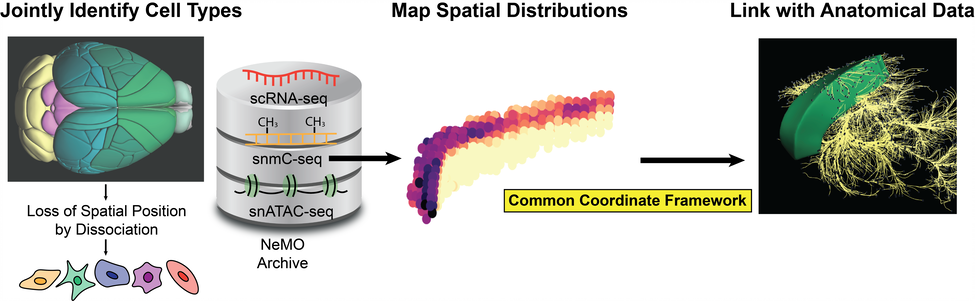
The brain contains diverse cell types that vary widely in characteristic properties and function in complex, interconnected circuits. A complete definition of cellular identity in the brain requires incorporating both molecular and anatomical properties, including gene expression, epigenetic regulation, spatial position and axonal projection patterns. However existing experimental approaches cannot measure all of these features simultaneously within individual cells. This inability to link anatomical and molecular features of cellular identity severely limits efforts to enumerate the brain's component cell types and assemble them into a circuit diagram. Computational integration of existing data provides a way to circumvent these challenges and construct a multi-modal atlas of brain cell types. Spatial transcriptomic measurements uniquely bridge imaging and molecular modalities, offering an opportunity to infer anatomical properties of cell types defined from single-cell transcriptome and epigenome data. The existence of whole-brain spatial transcriptomic data and projection data raises the exciting possibility of mapping dissociated single-cell data into space, thus linking molecular and anatomical features of neurons. We recently developed LIGER, an algorithm that can integrate multiple types of single-cell data to define brain cell types and recover the spatial positions of dissociated cells. Here, we will leverage our computational integration framework and use existing spatial transcriptomic data as a bridge to link disparate molecular and anatomical data. Specifically, we will: (1) apply LIGER to jointly define mouse brain cell types from single-cell transcriptomic and epigenomic data; (2) map this dissociated cell data in space using spatial transcriptomic data as a bridge; and (3) link cell types to anatomical properties using position within a common coordinate framework.
Project Leadership
Joshua Welch, Ph.D. (Principal Investigator)
Assistant Professor
Department of Computational Medicine and Bioinformatics
Department of Computer Science and Engineering
University of Michigan, Ann Arbor, MI
Project Data Types
This project will integrate existing datasets of the following types:
- Single-cell and single-nucleus RNA-seq, single-nucleus methylcytosine sequencing, and single-nucleus ATAC-seq data from adult mouse brain
- Spatial transcriptomic data from adult mouse brain
- Neuronal projection data from adult mouse
Related Resources
- Welch laboratory: https://welch-lab.github.io/
- LIGER software for integrating single-cell datasets: https://github.com/welch-lab/liger

