Accessible technologies for high-throughput, whole-brain reconstructions of molecularly characterized mammalian neurons
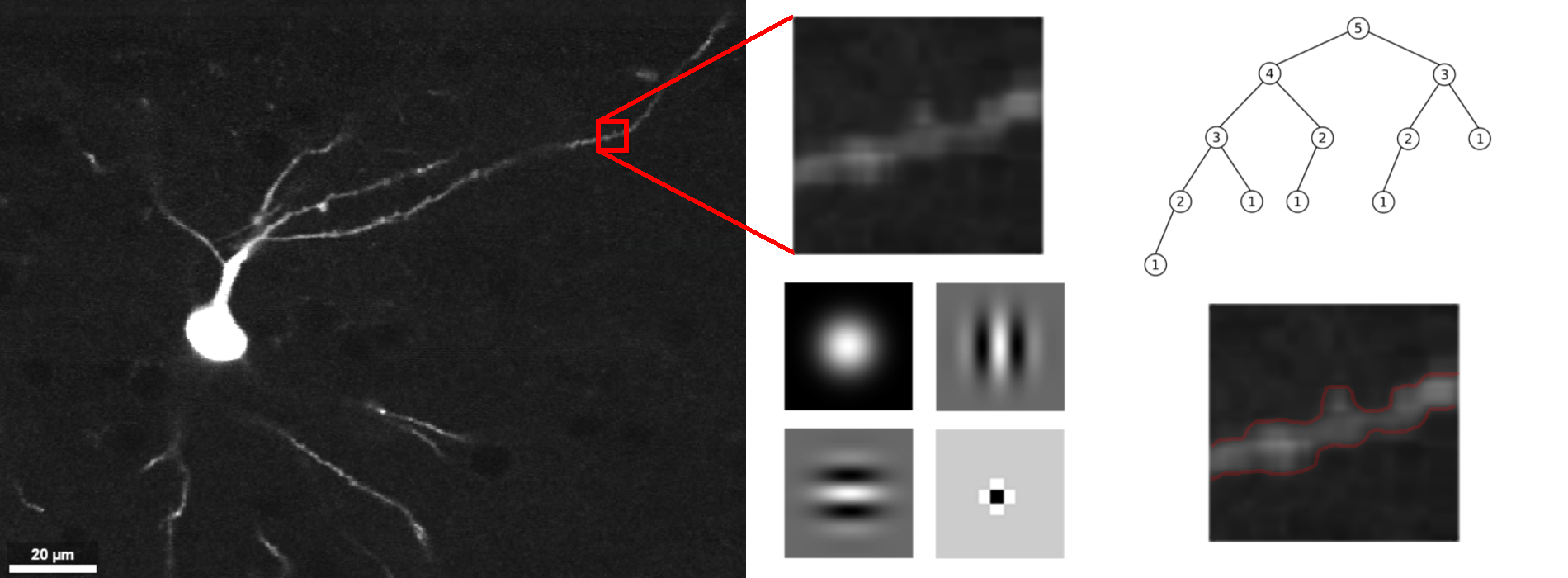
This project intends to automate neuron segmentation in sparsely labeled, sub-micron resolution images of complete mammalian brains. The terabyte scale of the data necessitates the use of efficient image processing, principled dimensionality reduction, and distributed computing. Algorithms will first be developed on existing two-photon microscopy images from HHMI Janelia, then extended to new experiments. The resulting open-source software will streamline large-scale investigations into neuron morphology.
Modern neuroscience seeks to explain behavior, including sensation, cognition and movement, in terms of information processing in neural circuits. Defined cell types are the nodes of neural circuits. Enumerating and characterizing the brain’s cell types and their connectivity is therefore necessary to understand how circuits process information and how they change in brain disorders. The mammalian brain contains astronomic numbers of neurons (mouse, 108; human, 1011), distributed across ~1,000 brain areas, each containing multiple cell types (likely ranging from 1 – 100)1. It seems clear that classification of cell types will require integrative analysis of at least two data types at the single-cell level: (1) molecular signature (e.g., transcriptome), and (2) anatomy (e.g., location and connectivity).
The major goal of the ‘Brain Light’ project is to build on the expertise of researchers at JHMI/JHU and Janelia to develop efficient and scalable methods to reconstruct the complete morphologies of molecularly characterized neurons. Over the last seven years, the MouseLight project at Janelia has developed semi-automated methods to facilitate the efficient reconstruction of entire individual neurons, including their axonal arbors, based on light microscopy2. MouseLight has reconstructed more than 1,000 projection neurons from the motor cortex, thalamus, hippocampus, and hypothalamus (http://ml-neuronbrowser.janelia.org/). The MouseLight data set has already revealed key information about cell types and their connectivity22,3,4,5,6.
Our research will build on the expertise at JHMI/JHU in neuroscience and neuro-informatics and experience gained with MouseLight to develop methodologies necessary to elevate the morphological reconstruction of molecularly defined neurons from an artisanal method to a technology that is scalable and suitable for the analysis of all neuron types across the mouse brain and, eventually, other mammalian brains. To achieve this goal, we will develop technology for imaging and reconstructing axon arborizations of individual molecularly defined neurons, thereby providing critical information about neuronal structure and connectivity. The data and methods will be made widely available to enable a community-wide effort to map neuronal connectivity at unprecedented scales and depth.
References:
- Bota, M. & Swanson, L. W. The neuron classification problem. Brain Res Rev 56, 79–88 (2007).
- Winnubst, J. et al. Reconstruction of 1,000 projection neurons reveals new cell types and organization of long-range connectivity in the mouse brain. bioRxiv 537233 (2019). doi:10.1101/537233
- Economo, M. N. et al. Distinct descending motor cortex pathways and their roles in movement. Nature 563, 79 (2018).
- Economo, M. N. et al. A platform for brain-wide imaging and reconstruction of individual neurons. eLife 5, e10566 (2016).
- Cembrowski, M. S. et al. Dissociable Structural and Functional Hippocampal Outputs via Distinct Subiculum Cell Classes. Cell 173, 1280-1292.e18 (2018).
- Hooks, B. M. et al. Topographic precision in sensory and motor corticostriatal projections varies across cell type and cortical area. Nat. Commun. 9, 3549 (2018).
Project Leadership
Ulrich Mueller, Ph.D. (Principal Investigator)
Bloomberg Distinguished Professor of Neuroscience and Biology
Johns Hopkins School of Medicine
http://neuroscience.jhu.edu/research/faculty/129
Micheal I. Miller, Ph.D. (Co-Investigator)
Bessie Darling Massey Professor and Director of Biomedical Engineering
Johns Hopkins University
https://engineering.jhu.edu/ece/faculty/miller-michael/
Dwight Bergles, Ph.D. (Co-Investigator)
Professor of Neuroscience and Otolaryngology-head & neck surgery
Johns Hopkins School of Medicine
http://neuroscience.jhu.edu/research/faculty/6
Solange Brown, M.D., Ph.D. (Co-Investigator)
Associate Professor of Neuroscience
Johns Hopkins School of Medicine
http://neuroscience.jhu.edu/research/faculty/11
Jayaram Chandrashekar, Ph.D. (Co-Investigator)
Scientific Operations Manager
Janelia Research Campus
https://www.janelia.org/people/jayaram-chandrashekar
Jeremiah Cohen, Ph.D. (Co-Investigator)
Assistant Professor of Neuroscience
Johns Hopkins School of Medicine
http://neuroscience.jhu.edu/research/faculty/15
Joshua Dudman, Ph.D. (Co-Investigator)
Senior Group Leader
Janelia Research Campus
https://www.janelia.org/people/joshua-dudman
Jan Funke, Ph.D. (Co-Investigator)
Group Leader
Janelia Research Campus
https://www.janelia.org/people/jan-funke
Don Geman, Ph.D. (Co-Investigator)
Professor Applied Math and Statistics
Johns Hopkins University
http://cis.jhu.edu/faculty/dgeman.php
Adam Hantman, Ph.D. (Co-Investigator)
Group Leader
Janelia Research Campus
https://www.janelia.org/people/adam-hantman
Natasha K. Hussain, Ph.D. (Collaborator)
Scientific Director, Kavli Neuroscience Discovery Institute
Johns Hopkins School of Medicine
https://kavlijhu.org/about/members/2
Alex Kolodkin, Ph.D. (Co-Investigator)
Professor of Neuroscience
Johns Hopkins School of Medicine
http://neuroscience.jhu.edu/research/faculty/44
Dan O’Connor, Ph.D. (Co-Investigator)
Associate Professor of Neuroscience
Johns Hopkins School of Medicine
http://neuroscience.jhu.edu/research/faculty/65
Tilak Ratnanather, Ph.D. (Co-Investigator)
Associate Research Professor
Johns Hopkins University
http://cis.jhu.edu/faculty/ratnan.php
Marshall Hussain Shuler, Ph.D. (Co-Investigator)
Associate Professor of Neuroscience
Johns Hopkins School of Medicine
http://neuroscience.jhu.edu/research/faculty/82
Nelson Spruston, Ph.D. (Co-Investigator)
Sr. Director, Scientific Programs
Janelia Research Campus
https://www.janelia.org/people/nelson-spruston
Scott Sternson, Ph.D. (Co-Investigator)
Group Leader
Janelia Research Campus
https://www.janelia.org/people/scott-sternson
Jeremias Sulam, Ph.D. (Co-Investigator)
Assistant Professor Biomedical Engineering
Johns Hopkins University
https://sites.google.com/view/jsulam
Karel Svoboda, Ph.D. (Co-Investigator)
Senior Group Leader
Janelia Research Campus
https://www.janelia.org/people/karel-svoboda
Joshua Vogelstein, Ph.D. (Co-Investigator)
Assistant Professor of Biomedical Engineering
Johns Hopkins University
https://www.bme.jhu.edu/faculty_staff/joshua-t-vogelstein-phd/
Project Data Types
- Multiplexed fluorescence in situ hybridization (FISH) for molecular characterization of sparsely labeled neurons
- Matched anatomical connectivity information by viral retrograde and Cre-dependent anterograde tracing and microscopy
- Brain clearing (CLARITY) facilitated morphology data
- high-contrast, high-resolution imaging of neuronal structure using serial block-face, two-photon microscopy (SBF2P) and whole-brain light sheet microscopy (WBLSM)
- Development of a cloud-compliant elastic data management platform to store and process terabyte-scale image volumes, by combining the functionality of the feature-rich local MouseLight workstation, and the commercial-cloud based NeuroDataCloud.
- Development of machine learning tools, including convolutional, deep learning, and action recognition, to overcome bottlenecks in the reconstruction speed of neuronal morphology.
- Multiphoton and lightsheet microscopy image data
- Apply computational methods (including Large Deformation Diffeomorphic Metric Mapping (LDDMM)) for more rapid and robust nonlinear registration of neurons
- Sparse labeling technologies and apply both SBF2P and WBLSM for morphological reconstruction and we will in a reiterative way molecularly characterize the molecular phenotype of thousands of the reconstructed neurons by post hoc multiplex FISH
Related Resources
- FreeSurfer suite: https://surfer.nmr.mgh.harvard.edu/fswiki; for neuroimaging analysis algorithms mapping histological information to in vivo studies.
- Mouselight: https://www.janelia.org/project-team/mouselight; database and viewer for neuroimaging analysis
- Neurodata cloud: https://neurodata.io/nd_cloud/; pipeline and protocols that make these large neuroscience datasets available to everyone
- HHMI Janelia: https://www.janelia.org/project-team/mouselight; providing state of the art images and their current processing pipeline
- CloudVolume: https://github.com/seung-lab/cloud-volume; convert image data into Neuroglancer Precomputed Format
- Amazon Web Services: https://aws.amazon.com/; cloud storage and computation, especially S3 (storage) and EC2 (computation)
- Neuroglancer: https://github.com/neurodata/neuroglancer; volumetric data viewer
- Python: https://www.python.org/; computational programming and popular libraries such as: numpy, scipy, skimage, matplotlib
- TensorFlow: https://www.tensorflow.org/; to use and adapt state of the art neuron segmentation algorithms

